The LIFE MedCLIFFS project addresses the management of Invasive Alien Plant Species (IAPS), which are an increasingly serious problem in the Mediterranean coast, closely linked to climate change, globalization and trade in ornamental plants. We are particularly interested in the invasive species of the Habitat of Community Interest 1240 – Vegetated sea cliffs of the Mediterranean coasts with endemic Limonium spp. (HCI1240) of the Natura 2000 network of the Costa Brava coast (NE Iberian Peninsula). The main objectives of the project are: (1) to enable participative networks for the early detection and rapid eradication by engaging volunteers with different levels of expertise in monitoring the distribution, occupancy and population abundance of IAPS; (2) to improve the rapid response systems by implementing an automatic risk assessment system specifically adapted to the HCI1240 and Costa Brava conditions that will include data obtained by citizen science and will be incorporated by competent administratives for their management structure to guide decisions about where to target surveillance and control activities; (3) to eradicate, control and contain the most severe IAPS, especially Carpobrotus spp., Opuntia spp. and Gazania rigens; and (4) to prevent the introduction of IAPS and to integrate biodiversity into market by developing a framework (with the collaboration of nurseries, gardening companies, local governments and the general public) that regulate the marketing and use of some ornamental plants potentially dangerous to biodiversity if they become invasive (and how they can be prevented).
LIFE medCLIFFS is lead by the Spanish Research Council (CSIC) through the Botanical Institute of Barcelona (IBB), and counts with five partners: the Government of Catalonia (GENCAT), the Provincial Council of Girona (DdGI), the Museum of Natural Science of Barcelona (MCNB), the Girona Association of Nurseries (AVG) and Flora Catalana association (FC). All of them are committed to sustain the project’s results in future regarding the prevention of the introduction pathways of ornamental plants, the early warning networks, and the control/eradication of the most aggressive IAPS in the project’s areas.
Invasive Alien Species (IAS) have become very important components of the floras and faunas across the world, being regarded as one of the main drivers of native species extinctions. Understanding the biological processes that underlie invasion is critical to effectively manage current invaders and to prevent future invasions. Among the possible evolutionary pathways leading to invasiveness, hybridisation is one of the most interesting and least well understood. Certainly, some of the most dangerous IAS are the product of hybridisation processes, highlighting the central role of this evolutionary mechanism in invasion success. Hybridisation has proven to contribute to the success of invasiveness thanks to increased fitness or adaptive evolution (in most cases, to new ecological niches), which can be the result of several genetic and genomic processes.
In recent years, some studies have already reported the role played by hybridisation in providing a larger pool of genetic diversity, counteracting the disadvantages of strong bottlenecks linked to the invasion process. In contrast, the potential effect of structural variation at the genomic level in the rapid adaptation of hybrid taxa is a promising but still understudied field of research. The main hypothesis of our project proposal (GENNIAlien) is that genomic structural changes in hybrids such as changes in ploidy level, chromosomal rearrangements or expansion/reduction of transposable elements (TE) drive the appearance of additional evolutionary novelty and diversity, leading to an enhanced invasion capacity. As models to test this hypothesis, we are using two widespread species systems that invade Mediterranean ecosystems: the Carpobrotus edulis-acinaciformis and the Kalanchoe × houghtonii complexes. Both groups represent contrasting examples of homoploid and polyploid hybridisation, where previous evidence already indicates that genomic structural variation is contributing to their invasion capacity.
By applying the latest methodological approaches to explore landscape genomics and niche dynamics in these two species systems, we will be able: (i) to disentangle the number and timing of introductions, as well as to track their history of hybridisation events; (ii) to test whether the effects of hybridisation on the genomic structure of individuals including changes in ploidy level, chromosome rearrangements and TE dynamism contribute to the evolution of invasiveness; and (iii) to see whether genetic and genomic changes in hybrid invasive plants imply niche shifts, as well as to unravel how these would affect invasion success under several scenarios of climate change. In this sense, GENNiAlien will expand the frontiers of knowledge of invasion biology and evolutionary biology, two fields of study that are increasingly important in the context of the current climate and biodiversity crisis.
Funding agency:
Ministry of Science and Innovation of Spain
Reference:
PID2020-119163GB-I00
Principal researcher:
Drs. Jordi López-Pujol & Sònia Garcia
Duration:
2021–2025
Amount funded: 109,021 €
Number of researchers:
4 (plus a FPI PhD student)

Reports on climate change (CC) in mountain areas highlight the vulnerability of Pyrenees, particularly in terms of biodiversity, the lack of information on the effects of CC and the difficulty of identifying factors within global changes. The Pyrenean Climate Change Observatory (OPCC) is a cross-border initiative of territorial cooperation that aims to monitor and understand the effects of CC in the Pyrenees for improving its adaptation. The project FLORAPYR-AVANCE establishes a cross-border agreement with research centres of Andorra, Spain and France with the objective of contributing to the OPCC and combating the vulnerability of this flora so rich in endemism and so sensitive to increases in temperature. The previous project FLORAPYR (06/2016-06/2019, http://www.florapyr.opcc-ctp.org/) compiled information about the flora in Pyrenees in the web page of FLORAPYR.
Related to the alien plants, our team takes part in this project through two actions: (1) developing a citizen science monitoring of invasive flora and (2) improving the knowledge of alien flora in the Pyrenees. Taking advantage of the information compiled in the previous FLORAPYR, in this project we will elaborate reports of monitoring protocols of alien plants, including maps of these plants in Pyrenees for inclusion as an information layer in the OPCC web portal. The coordination of volunteers will permit the detection of non-native plants in some areas in the south-eastern Pyrenees. The final objective is to compile a checklist of alien species in Pyrenees, which is the first step for revealing the patterns of biological invasions and developing eradication strategies from a common perspective in the whole Pyrenees.
Funding agency: Interreg Poctefa,
program of the European Union
Reference: EFA322/19
Principal researcher: Dr. Gérard Largier

This project is aimed at generating basic and applied information on the status of alien species that have been introduced into mainland Ecuador (EC). The project will help in understanding the causes of biological invasions in EC, as a tool for the prevention of future invasions. Additionally, it would allow identifying and quantifying the consequences that these species can generate on natural resources, the economy and human health.
The project has three approaches. The first approach is to obtain an inventory of alien species in EC protected areas (PAs) and to assess which characteristics of PAs (spatial and anthropogenic) may increase the vulnerability of these areas to the establishment of alien species. The second approach is to generate information on distribution to analyze the causes and consequences of the establishment of the American bullfrog (Lithobates catesbeianus), in EC, an amphibian included in the list of the 100 most harmful invasive species in the world. The third approach will allow us to understand through a case study (genus Kalanchoe) how the introduction of plant species for ornamental and medicinal purposes can cause invasions in natural habitats.
Financial agency: Universidad Espíritu Santo (Ecuador)
Reference: S/N
Principal Researcher: Dr. Ileana Herrera (Escuela de Ciencias Ambientales, Universidad Espíritu Santo)
Duration: 2020-2022
Amount funded: 33,190.94 USD
Number of researchers: 12
Hemp (Cannabis sativa and related taxa and races) and, especially, Indian cannabis (name of one of its most used races as medicinal or stupefacient), is one of the most well-known, popular and used plants in the world. This notwithstanding, its integral study (starting with its taxonomy and until its present and potential applications) is far from being complete. In this project we foresee for the first time a multidisciplinary study linking the fields of morphology, systematics, genetics (including cytogenetics), ethnobotany, phytochemistry and pharmacology, and this concerning one of the species with the biggest biocultural relevance and potential in health in its largest sense, among other aspects.
We will use innovative techniques in morphometry, directed ethnobotanical interviews, genotyping and other genetic techniques, chemical analysis of phytocannabinoids and terpenes, and pharmacological investigations in cancer cell lines. The obtained results will allow us establishing in detail the taxonomy of the genus and its origin, and valuing in a related manner the diversity and variability in its morphometric characters, its genetic structure, its chemical composition, its traditional uses and management, and its medicinal potential (and, secondarily, others such as food and textile). In addition, the project will create a germplasm bank with seeds of all studied populations, which will remain available for possible future uses. The formation and the experience in different areas of the investigator and work teams ensure us to be able to accept the challenge with guarantees. The other big pillar in which the strength of the project is sustained is sampling, reaching the areas where hemp has its centres of origin and domestication, and wild occurrence. The field trips foreseen, mostly, and, to a lesser extent, the contributions from collaborators, together with the access to germplasm of cultivated races, will allow us to study the diversity of Cannabis as it has never been performed to date, at a large and detailed populational level, with an extensive prospection area and with very different approaches which will be considered jointly. This study will constitute an innovative contribution to several objectives included in the societal challenges, such as the efficiency and the use of natural resources and raw materials in a climate change scenario, and population health and wellness in a demographic change one. Additionally, the obtained results will be the basis for future Cannabis applications in medicine and in other fields.
Funding agency: State R&D&I program of Spain
(Ministerio de Economía, Industria y Competitividad)
Reference: CGL2017-84297-R
Principal researcher: Dra. Teresa Garnatje
Duration: 2018 – 2021
Amount funded: 217,800 €
Number of researchers: 12

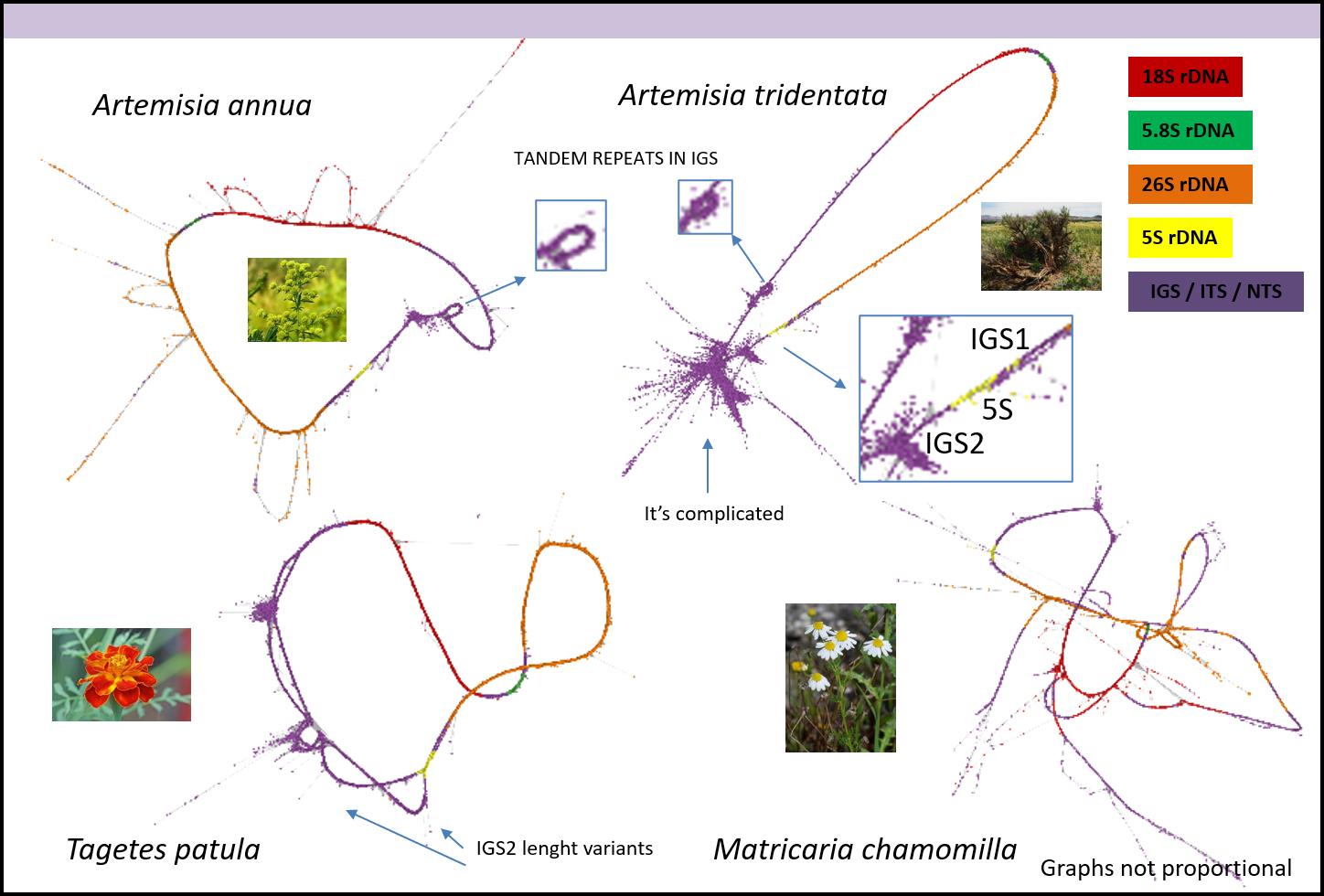
rDNA-Life aims to explore the evolutionary dynamism of ribosomal RNA genes (rDNA), a dynamism that is surprisingly high given their crucial role in the synthesis of ribosomes and thus for life-on-Earth. There are four main rDNAs: the 18S, 5.8S, and 26S rRNA genes, which are coded in a single operon, and the 5S rRNA gene usually coded outside that operon. These rDNAs are present in high numbers, from 50 to 13,000 copies per cell, and typically tandemly arranged. However, despite their importance and abundance, several aspects of the biology of these critical molecules for life remain enigmatic, particularly their mode of molecular evolution, genomic arrangements and epigenetic regulation. rDNA-Life aims to deconstruct ribosomal DNA with the purposes of: (1) defining the model of evolution that best explains rDNA’s high dynamism and the differences depending on the rDNA type, (2) assessing the impact of epigenetic processes, like methylation, on rDNA evolution and (3) establishing the landscape of rDNA arrangements across the Tree-of-Life.
TrDNA-Life’s approach is mainly focused in the plant kingdom, and we will use as models for our research two species pairs from genera Artemisia and Tragopogon. Their evolutionary relatedness (both belong to family Asteraceae) coupled with contrasting rDNA arrangements (the former genus with all rRNA genes linked, the latter with 5S rDNA separated from the rest), different genome sizes, ploidy levels, and numbers and positions of rDNA loci in chromosomes make them compelling to study, and their mutual comparison promises insightful results.
With next-generation sequencing (NGS) and advances in epigenetics, we will address central questions that hitherto would have been impossible. We will study plant ribosomal RNA genes (rDNA) to gain novel, fundamental insights into genomic mechanisms such as concerted evolution, pseudogenisation, genome size control and genome stability, and about the relationship between transposable elements and rDNA. We anticipate results of broad interest to evolutionary biologists with impact in our understanding of molecular evolution and repetitive DNA organisation. rDNA-Life will generate fundamental knowledge on the mechanisms involved in the maintenance of rDNA stability, including its expression and epigenetic regulation.
The multidisciplinary approach of rDNA-Life combines botany and evolutionary biology with genomics and epigenetics and the use of state-of-the-art techniques, including NGS, whole genome bisulfite sequencing (WGBS) and bioinformatics approaches to analyse sequence data. rDNA-Life also involves the use of nanobiotechnological methods to produce genome maps and confocal microscopy to observe methylation status of rDNA in extended DNA fibers.
The project’s feasibility is ensured by the strong background in rDNA biology of Dr. Sònia Garcia (principal investigator, PI) and the members of the working team. Several of the researchers involved in rDNA-Life collaborate with the establishment and curation of the Plant rDNA database, an underpinning resource that will be enhanced and utilised in the project’s framework. Besides, rDNA-Life will certainly promote an independent leadership position in science of the PI. Finally, while exploring core questions in the research interests of the PI’s institution (Institut Botànic de Barcelona, IBB, CSIC-Ajuntament de Barcelona), rDNA-Life will raise national and international excellence in evolutionary biology.
KEY WORDS: bioinformatics, chromosome, epigenetics, genome organisation, genome size, molecular evolution, NGS, repetitive DNA, ribosomal DNA, transposable elements
Funding agency: State R&D&I program of Spain
(Ministerio de Economía y Competitividad)
Reference: CGL2016-75694-P
Principal researcher: Dra. Sònia Garcia
Duration: 2016 – 2020
Amount funded: 160,930 €
Number of researchers: 1